NeuThera - AI-Driven Drug Discovery Toolkit
Inspiration
Traditional drug discovery methods are hindered by high costs, long timelines upwards of 20 years, and a high attrition rate in clinical trials. With the rise of AI and computational biology -- Generative models and in-silico validation techniques have shown promise in accelerating early-stage drug design. However, current models often operate independently of each other, lacking an integrated pipeline for end-to-end drug candidate generation.
NeuThera was proposed as a modular drug discovery starter toolkit, integrating multiple state-of-the-art generative molecular design models such as TamGen, chemBERTa, and DeepPurpose with other advanced tooling like graph-based retrieval, molecular fingerprint embeddings, ADMET profiling, etc.
What it does
The goal is to create a fully modular agentic AI framework capable of generating, validating, and optimizing novel drug candidates in an end-to-end pipeline - creating and maintaining a graph database of all FDA approved and generated compounds for further repurposing or discovery.
NeuThera supports both end-to-end drug discovery automation and interactive, user-guided drug design workflows.
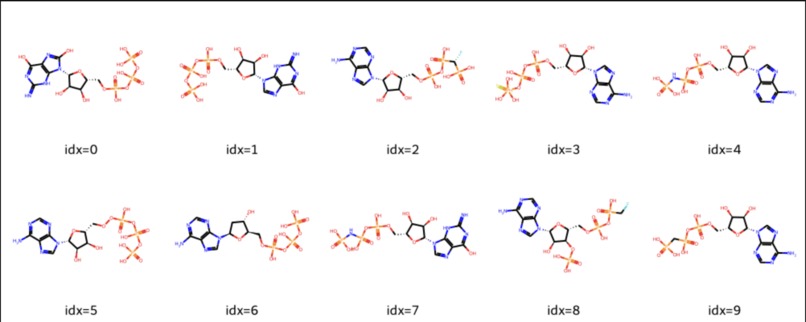
- Generate de novo molecules: using multiple SOTA generative models, including Transformer-based molecular generator models. Ability to generate millions of compounds with complete virtual ADMET profiling for validation. All generated compounds are viable in the real world.
- biomedical knowledge-base: Combines 15+ datasets from BioSNAP, DrugBank, Chembl, and PDB to create a starting point for drug discovery and repurposing.
- Molecular Vector Embedding Space: Vector embeddings from chemBERTa facilitate lead optimization, scaffold hopping, and similarity search.
- In-Silico Testing: With generated compounds and target proteins, uses DeepPurpose to predict binding affinity without requiring real world intervention
How we built it
NeuThera is architected as a multi-model system, integrating specialized tools and frameworks to create a starter toolkit for end-to-end drug discovery. The modular nature of the technology used allows researchers to expand the tooling, as all necessary data and functions are already provided.
Core Modules
- TamGen: A transformer-based generative model for de novo molecular design, trained on 20 million+ SMILES of known bioactive compounds. All generated compounds are validated and docked on target protein using DockBox before final result.
- DeepPurpose: A multi-model framework used to check the binding affinity between a drug and a target protein. As all generated compounds from TamGen are already validated with docking coordinates, DeepPurpose fits in perfecting with this pipeline.
- chemBERTa: Transformer-based model trained on 120+ million SMILES to create meaningful vector embeddings. Used to create vector space of all FDA Approved drugs + generated compounds for advanced analytics.
Knowledge Graph & Retrieval Modules
- ArangoDB: A multimodal database storing structured relationships between genes, proteins, diseases, drugs, and molecular structures.
- Graph-based querying for retrieving relevant molecular information, linking known drug-target interactions to novel compounds.
- ArangoDB: A multimodal database storing structured relationships between genes, proteins, diseases, drugs, and molecular structures.
Molecular Similarity & Lead Optimization
- Morgan/Circular Fingerprints: Used for molecular embedding, leveraging Tanimoto similarity and vector representations.
- Vector Space Search: for identifying structurally similar compounds within large-scale chemical libraries.
- Graph-based lead optimization: Using molecular graphs and chemical property analysis to refine generated compounds.
- Morgan/Circular Fingerprints: Used for molecular embedding, leveraging Tanimoto similarity and vector representations.
GraphRAG & Biomedical Reasoning
- GraphRAG framework Augments traditional RAG (Retrieval-Augmented Generation) methods by incorporating graph-based reasoning for biomedical data
- Ontological mapping: Integrates structured biomedical ontologies (e.g., DrugBank, ChEMBL) to enhance retrieval and inference.
Challenges we ran into
- Data inconsistencies and scattered datasets: As the biomedical data space is vast and varied from field to field (Most use different identifiers), it required us multiple days to put together the graph database using only open-source mappings. A lot of the datasets used required combining multiple different mappings to get the required fields.
- Outdated libraries and models: As most of the technology we're using is state of the art, they are not mature enough to work well with different technologies. Example: TamGen only has around 30 stars on github and required very specific library versions to work. We had to rewrite some of the code to work for our needs and combine with our tech stack. Sadly, we've had to discard nx-arangodb as it does not function well with TamGen which is a core unit of our project.
- Scaling the solution: As college students with almost no resources, we've had to keep our AI model (GPT-4o) usage to a bare-minimum. As such, we could not fine tune it to work as we intended, though it works well ~75% of the time. We wish to work on this project for a long and with adequate funding, we can run a model with much better reasoning abilities and fine tune to be extremely useful.
Accomplishments that we're proud of
- TamGen code rewrite to work without CUDA: Our code rewrite for TamGen can be a valid contribution for their repository.
- Biomedical knowledge retrieval: Combing 15+ datasets enables us to do structured hypothesis-driven drug design.
- Scalable molecular similarity search: using vector embeddings for rapid lead identification and generated drug repurposing.
What we learned
- Got a glimpse of how cutting edge research for generative AI works.
- How vast and wonderful the open source community for biomedical fields are.
- Working with multiple tooling and creating coherence between them.
- Using SOTA technologies like ArangoDB and GraphRAG to create meaningful toolings
What's Next for NeuThera?
Enhancing generative models with a feedback loop:
- Integrating diffusion-based generative modeling to improve the novelty, drug-likeness, and synthetic accessibility of generated compounds.
- Implementing a reinforcement-driven feedback loop for iterative optimization
- TamGen generates an initial batch of 10000 de novo compounds.
- DeepPurpose predicts binding affinity for each compound with the target protein.
- Compound encoders select the highest-scoring compounds as reference for next batch.
- Loop continues until convergence, maximizing binding affinity while optimizing ADMET properties.
- TamGen generates an initial batch of 10000 de novo compounds.
- Integrating diffusion-based generative modeling to improve the novelty, drug-likeness, and synthetic accessibility of generated compounds.
Automated lead optimization and drug repurposing:
- Integrating Graph Neural Networks (GNNs) on molecular embeddings to predict novel indications for existing compounds.
- Scaffold hopping via vector space traversal to explore chemically diverse, high-affinity derivatives.
- Integrating Graph Neural Networks (GNNs) on molecular embeddings to predict novel indications for existing compounds.
Modular tooling for user-defined extensions:
- Allowing users to integrate custom generative models, scoring algorithms, functions, etc.
- Supporting custom scoring functions, including multi-objective optimization for ADMET, pharmacokinetics, and toxicity filtering.
- Allowing users to integrate custom generative models, scoring algorithms, functions, etc.
Advanced UI/UX for visualization and interactive drug design:
- Developing an interactive web platform with:
- AI-Chatbot system
- Molecular visualization for ligand-protein interactions and generated compounds.
- Graph-based relationship exploration to analyze drug-target-disease links.
- Live compound generation and scoring dashboard, enabling real-time feedback on molecular design.
- Developing an interactive web platform with:









Log in or sign up for Devpost to join the conversation.