Abstract
Purpose
To validate five automated structural MRI quality assessment tools against expert visual ratings and assess their reliability, validity, and practical utility for large-scale neuroimaging research.
Methods
Structural MRI data from 92 participants (ages 5–20 years) in the Healthy Brain Network were analyzed. Five tools—FreeSurfer, FSQC, MRIQC, BrainSuite, and the Computational Anatomy Toolbox (CAT)—were evaluated for computational reproducibility, convergent validity with expert ratings, and discriminative ability between expert-rated “Pass” and “Fail” scans. Expert ratings served as the reference standard.
Results
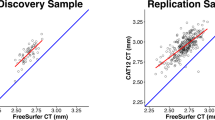
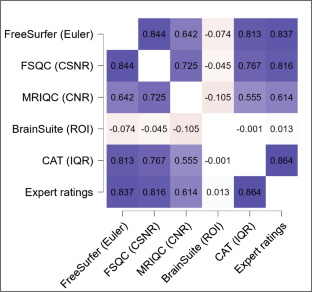
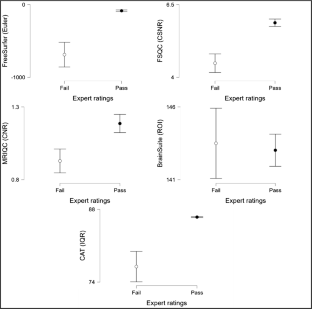
All tools demonstrated excellent computational reproducibility. FreeSurfer, FSQC, MRIQC, and CAT correlated strongly with expert ratings and discriminated effectively between “Pass” and “Fail” scans. FreeSurfer, FSQC, and CAT achieved near-perfect classification accuracy, although CAT systematically assigned higher scores even to poor-quality scans, suggesting the need for stricter thresholds. MRIQC aligned less strongly but captured complementary quality dimensions. BrainSuite metrics did not correspond to expert ratings or separate scan quality.
Conclusion
Automated MRI quality assessment tools provide reliable and scalable alternatives to manual inspection. FreeSurfer, FSQC, and CAT approach expert-level accuracy but require careful calibration, while MRIQC provides complementary insights despite weaker alignment. Adoption of automated approaches, with awareness of tool-specific limitations, can enhance reproducibility, efficiency, and rigor in large-scale neuroimaging studies.
Similar content being viewed by others
Data availability
We report how we determined our sample size, all data exclusions, all manipulations, and all measures in the study, and the study follows Journal Article Reporting Standards [71]. All data, analysis code, and research materials are available at https://osf.io/ypfd2/, and are shared under a Creative Commons Attribution 4.0 International (CC BY 4.0) license. For additional information about the data and research materials, please visit https://fcon_1000.projects.nitrc.org/indi/cmi_healthy_brain_network/MRI_EEG.html. This study’s design and its analysis were preregistered at https://osf.io/9s4g7.
References
Mills KL, Tamnes CK (2014) Methods and considerations for longitudinal structural brain imaging analysis across development. Dev Cogn Neurosci 9:172–190. https://doi.org/10.1016/j.dcn.2014.04.004
Poldrack RA (2012) The future of fMRI in cognitive neuroscience. Neuroimage 62(2):1216–1220. https://doi.org/10.1016/j.neuroimage.2011.08.007
Alexander-Bloch A, Clasen L, Stockman M, Ronan L, Lalonde F, Giedd J, Raznahan A (2016) Subtle in-scanner motion biases automated measurement of brain anatomy from in vivo MRI. Hum Brain Mapp 37(7):2385–2397. https://doi.org/10.1002/hbm.23180
Gilmore AD, Buser NJ, Hanson JL (2021) Variations in structural MRI quality significantly impact commonly used measures of brain anatomy. Brain Inform 8:7. https://doi.org/10.1186/s40708-021-00128-2
Reuter M, Tisdall MD, Qureshi A, Buckner RL, van der Kouwe AJW, Fischl B (2015) Head motion during MRI acquisition reduces gray matter volume and thickness estimates. Neuroimage 107:107–115. https://doi.org/10.1016/j.neuroimage.2014.12.006
Dosenbach NUF, Koller JM, Earl EA, Miranda-Dominguez O, Klein RL, Van AN, Snyder AZ, Nagel BJ, Nigg JT, Nguyen AL, Wesevich V, Greene DJ, Fair DA (2017) Real-time motion analytics during brain MRI improve data quality and reduce costs. Neuroimage 161:80–93. https://doi.org/10.1016/j.neuroimage.2017.08.025
Greene DJ, Koller JM, Hampton JM, Wesevich V, Van AN, Nguyen AL, Hoyt CR, McIntyre L, Earl EA, Klein RL, Shimony JS, Petersen SE, Schlaggar BL, Fair DA, Dosenbach NUF (2018) Behavioral interventions for reducing head motion during MRI scans in children. Neuroimage 171:234–245. https://doi.org/10.1016/j.neuroimage.2018.01.023
Alexander LM, Escalera J, Ai L, Andreotti C, Febre K, Mangone A, Vega-Potler N, Langer N, Alexander A, Kovacs M, Litke S, O’Hagan B, Andersen J, Bronstein B, Bui A, Bushey M, Butler H, Castagna V, Camacho N, Chan E, Citera D, Clucas J, Cohen S, Dufek S, Eaves M, Fradera B, Gardner J, Grant-Villegas N, Green G, Gregory C, Hart E, Harris S, Horton M, Kahn D, Kabotyanski K, Karmel B, Kelly SP, Kleinman K, Koo B, Kramer E, Lennon E, Lord C, Mantello G, Margolis A, Merikangas KR, Milham J, Minniti G, Neuhaus R, Levine A, Osman Y, Parra LC, Pugh KR, Racanello A, Restrepo A, Saltzman T, Septimus B, Tobe R, Waltz R, Williams A, Yeo A, Castellanos FX, Klein A, Paus T, Leventhal BL, Craddock RC, Koplewicz HS, Milham MP (2017) An open resource for transdiagnostic research in pediatric mental health and learning disorders. Sci Data 4(1):170181. https://doi.org/10.1038/sdata.2017.181
Shafiei G, Esper NB, Hoffmann MS, Ai L, Chen AA, Cluce J, Covitz S, Giavasis S, Lane C, Mehta K, Moore TM, Salo T, Tapera TM, Calkins ME, Colcombe S, Davatzikos C, Gur RE, Gur RC, Pan PM, Satterthwaite TD (2025) Reproducible brain charts: an open data resource for mapping brain development and its associations with mental health. Neuron 113(22):3758-3779.e3756. https://doi.org/10.1016/j.neuron.2025.08.026
Elyounssi S, Kunitoki K, Clauss JA, Laurent E, Kane KA, Hughes DE, Hopkinson CE, Bazer O, Sussman RF, Doyle AE, Lee H, Tervo-Clemmens B, Eryilmaz H, Hirschtick RL, Barch DM, Satterthwaite TD, Dowling KF, Roffman JL (2025) Addressing artifactual bias in large, automated MRI analyses of brain development. Nat Neurosci 28(8):1787–1796. https://doi.org/10.1038/s41593-025-01990-7
Casey BJ, Cannonier T, Conley MI, Cohen AO, Barch DM, Heitzeg MM, Soules ME, Teslovich T, Dellarco DV, Garavan H, Orr CA, Wager TD, Banich MT, Speer NK, Sutherland MT, Riedel MC, Dick AS, Bjork JM, Thomas KM, Chaarani B, Mejia MH, Hagler DJ Jr., Daniela Cornejo M, Sicat CS, Harms MP, Dosenbach NUF, Rosenberg M, Earl E, Bartsch H, Watts R, Polimeni JR, Kuperman JM, Fair DA, Dale AM (2018) The Adolescent Brain Cognitive Development (ABCD) study: imaging acquisition across 21 sites. Dev Cogn Neurosci 32:43–54. https://doi.org/10.1016/j.dcn.2018.03.001
Marek S, Tervo-Clemmens B, Calabro FJ, Montez DF, Kay BP, Hatoum AS, Donohue MR, Foran W, Miller RL, Hendrickson TJ, Malone SM, Kandala S, Feczko E, Miranda-Dominguez O, Graham AM, Earl EA, Perrone AJ, Cordova M, Doyle O, Moore LA, Conan GM, Uriarte J, Snider K, Lynch BJ, Wilgenbusch JC, Pengo T, Tam A, Chen J, Newbold DJ, Zheng A, Seider NA, Van AN, Metoki A, Chauvin RJ, Laumann TO, Greene DJ, Petersen SE, Garavan H, Thompson WK, Nichols TE, Yeo BTT, Barch DM, Luna B, Fair DA, Dosenbach NUF (2022) Reproducible brain-wide association studies require thousands of individuals. Nature 603(7902):654–660. https://doi.org/10.1038/s41586-022-04492-9
Sonuga-Barke EJS (2023) Editorial: “safety in numbers”? Big data discovery strategies in neuro-developmental science – contributions and caveats. J Child Psychol Psychiatry 64(1):1–3. https://doi.org/10.1111/jcpp.13723
Spisak T, Bingel U, Wager TD (2023) Multivariate BWAS can be replicable with moderate sample sizes. Nature 615:E4–E7. https://doi.org/10.1038/s41586-023-05745-x
Esteban O, Birman D, Schaer M, Koyejo OO, Poldrack RA, Gorgolewski KJ (2017) MRIQC: advancing the automatic prediction of image quality in MRI from unseen sites. PLoS One 12(9):e0184661. https://doi.org/10.1371/journal.pone.0184661
Williams B, Hedger N, McNabb CB, Rossetti GMK, Christakou A (2023) Inter-rater reliability of functional MRI data quality control assessments: a standardised protocol and practical guide using pyfMRIqc. Front Neurosci 17:1070413. https://doi.org/10.3389/fnins.2023.1070413
Provins C, Savary É, Sanchez T, Mullier E, Barranco J, Fischi-Gómez E, Alemán-Gómez Y, Richiardi J, Poldrack RA, Hagmann P, Esteban O (2025) Removing facial features from structural MRI images biases visual quality assessment. PLoS Biol 23(4):e3003149. https://doi.org/10.1371/journal.pbio.3003149
Hendriks J, Mutsaerts H-J, Joules R, Peña-Nogales Ó, Rodrigues PR, Wolz R, Burchell GL, Barkhof F, Schrantee A (2024) A systematic review of (semi-)automatic quality control of T1-weighted MRI scans. Neuroradiology 66(1):31–42. https://doi.org/10.1007/s00234-023-03256-0
Bhalerao G, Gillis G, Dembele M, Suri S, Ebmeier K, Klein J, Hu M, Mackay C, Griffanti L (2025) Automated quality control of T1-weighted brain MRI scans for clinical research datasets: methods comparison and design of a quality prediction classifier. Imaging Neurosci 3:4. https://doi.org/10.1162/IMAG.a.4
Bedford SA, Ortiz-Rosa A, Schabdach JM, Costantino M, Tullo S, Piercy T, Lai M-C, Lombardo MV, Di Martino A, Devenyi GA, Chakravarty MM, Alexander-Bloch AF, Seidlitz J, Baron-Cohen S, Bethlehem RAI (2023) The impact of quality control on cortical morphometry comparisons in autism. Imaging Neurosci 1:1–21. https://doi.org/10.1162/imag_a_00022
Cosgrove KT, McDermott TJ, White EJ, Mosconi MW, Thompson WK, Paulus MP, Cardenas-Iniguez C, Aupperle RL (2022) Limits to the generalizability of resting-state functional magnetic resonance imaging studies of youth: an examination of ABCD Study® baseline data. Brain Imaging Behav 16:1919–1925. https://doi.org/10.1007/s11682-022-00665-2
Dale AM, Fischl B, Sereno MI (1999) Cortical surface-based analysis: I. segmentation and surface reconstruction. Neuroimage 9(2):179–194. https://doi.org/10.1006/nimg.1998.0395
Fischl B (2012) FreeSurfer. Neuroimage 62(2):774–781. https://doi.org/10.1016/j.neuroimage.2012.01.021
Kim Y, Joshi AA, Choi S, Joshi SH, Bhushan C, Varadarajan D, Haldar JP, Leahy RM, Shattuck DW (2024) BrainSuite BIDS app: containerized workflows for MRI analysis. bioRxiv. https://doi.org/10.1101/2023.03.14.532686
Gaser, C., Dahnke, R., Thompson, P. M., Kurth, F., Luders, E., & the Alzheimer’s Disease Neuroimaging Initiative. (2024). CAT: A computational anatomy toolbox for the analysis of structural MRI data. GigaScience, 13. https://doi.org/10.1093/gigascience/giae049
Han X, Jovicich J, Salat D, van der Kouwe A, Quinn B, Czanner S, Busa E, Pacheco J, Albert M, Killiany R, Maguire P, Rosas D, Makris N, Dale A, Dickerson B, Fischl B (2006) Reliability of MRI-derived measurements of human cerebral cortical thickness: the effects of field strength, scanner upgrade and manufacturer. Neuroimage 32(1):180–194. https://doi.org/10.1016/j.neuroimage.2006.02.051
Jovicich J, Czanner S, Greve D, Haley E, van der Kouwe A, Gollub R, Kennedy D, Schmitt F, Brown G, MacFall J, Fischl B, Dale A (2006) Reliability in multi-site structural MRI studies: effects of gradient non-linearity correction on phantom and human data. Neuroimage 30(2):436–443. https://doi.org/10.1016/j.neuroimage.2005.09.046
Takao H, Hayashi N, Ohtomo K (2011) Effect of scanner in longitudinal studies of brain volume changes. J Magn Reson Imaging 34(2):438–444. https://doi.org/10.1002/jmri.22636
Takao H, Hayashi N, Ohtomo K (2014) Effects of study design in multi-scanner voxel-based morphometry studies. Neuroimage 84:133–140. https://doi.org/10.1016/j.neuroimage.2013.08.046
Mugler JP III, Brookeman JR (1990) Three-dimensional magnetization-prepared rapid gradient-echo imaging (3D MP RAGE). Magn Reson Med 15(1):152–157. https://doi.org/10.1002/mrm.1910150117
Hoopes A, Mora JS, Dalca AV, Fischl B, Hoffmann M (2022) SynthStrip: skull-stripping for any brain image. Neuroimage 260:119474. https://doi.org/10.1016/j.neuroimage.2022.119474
Billot B, Greve DN, Puonti O, Thielscher A, Van Leemput K, Fischl B, Dalca AV, Iglesias JE (2023) SynthSeg: segmentation of brain MRI scans of any contrast and resolution without retraining. Med Image Anal 86:102789. https://doi.org/10.1016/j.media.2023.102789
Hoffmann M, Billot B, Greve DN, Iglesias JE, Fischl B, Dalca AV (2022) SynthMorph: learning contrast-invariant registration without acquired images. IEEE Trans Med Imaging 41(3):543–558. https://doi.org/10.1109/tmi.2021.3116879
Sled JG, Zijdenbos AP, Evans AC (1998) A nonparametric method for automatic correction of intensity nonuniformity in MRI data. IEEE Trans Med Imaging 17(1):87–97. https://doi.org/10.1109/42.668698
Fischl B, Liu A, Dale AM (2001) Automated manifold surgery: constructing geometrically accurate and topologically correct models of the human cerebral cortex. IEEE Trans Med Imaging 20(1):70–80. https://doi.org/10.1109/42.906426
Ségonne F, Pacheco J, Fischl B (2007) Geometrically accurate topology-correction of cortical surfaces using nonseparating loops. IEEE Trans Med Imaging 26(4):518–529. https://doi.org/10.1109/TMI.2006.887364
Fischl B, Dale AM (2000) Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc Natl Acad Sci U S A 97(20):11050–11055. https://doi.org/10.1073/pnas.200033797
Fischl B, Sereno MI, Dale AM (1999) Cortical surface-based analysis: II: inflation, flattening, and a surface-based coordinate system. Neuroimage 9(2):195–207. https://doi.org/10.1006/nimg.1998.0396
Destrieux C, Fischl B, Dale A, Halgren E (2010) Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. Neuroimage 53(1):1–15. https://doi.org/10.1016/j.neuroimage.2010.06.010
Rosen AFG, Roalf DR, Ruparel K, Blake J, Seelaus K, Villa LP, Ciric R, Cook PA, Davatzikos C, Elliott MA, de Garcia La Garza A, Gennatas ED, Quarmley M, Schmitt JE, Shinohara RT, Tisdall MD, Craddock RC, Gur RE, Gur RC, Satterthwaite TD (2018) Quantitative assessment of structural image quality. Neuroimage 169:407–418. https://doi.org/10.1016/j.neuroimage.2017.12.059
Reuter M, Wolter F-E, Peinecke N (2006) Laplace-Beltrami spectra as ‘shape-DNA’ of surfaces and solids. Comput Aided Des 38(4):342–366. https://doi.org/10.1016/j.cad.2005.10.011
Wachinger C, Golland P, Kremen W, Fischl B, Reuter M (2015) BrainPrint: a discriminative characterization of brain morphology. Neuroimage 109:232–248. https://doi.org/10.1016/j.neuroimage.2015.01.032
Gorgolewski, K., Burns, C. D., Madison, C., Clark, D., Halchenko, Y. O., Waskom, M. L., & Ghosh, S. S. (2011). Nipype: A Flexible, Lightweight and Extensible Neuroimaging Data Processing Framework in Python. Frontiers in Neuroinformatics, Volume 5. https://doi.org/10.3389/fninf.2011.00013
Jenkinson M, Beckmann CF, Behrens TEJ, Woolrich MW, Smith SM (2012) FSL. Neuroimage 62(2):782–790. https://doi.org/10.1016/j.neuroimage.2011.09.015
Avants BB, Tustison NJ, Song G, Cook PA, Klein A, Gee JC (2011) A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage 54(3):2033–2044. https://doi.org/10.1016/j.neuroimage.2010.09.025
Cox RW, Hyde JS (1997) Software tools for analysis and visualization of fMRI data. NMR Biomed 10(4–5):171–178
Cabalo DG, Leppert IR, Thevakumaran R, DeKraker J, Hwang Y, Royer J, Kebets V, Tavakol S, Wang Y, Zhou Y, Benkarim O, Eichert N, Paquola C, Doyon J, Tardif CL, Rudko D, Smallwood J, Rodriguez-Cruces R, Bernhardt BC (2025) Multimodal precision MRI of the individual human brain at ultra-high fields. Sci Data 12(1):526. https://doi.org/10.1038/s41597-025-04863-7
Wang B, Zhang X, Zhang L, Kong X-Z (2025) A naturalistic fMRI dataset in response to public speaking. Sci Data 12(1):659. https://doi.org/10.1038/s41597-025-05017-5
Shattuck DW, Leahy RM (2002) BrainSuite: an automated cortical surface identification tool. Med Image Anal 6(2):129–142. https://doi.org/10.1016/S1361-8415(02)00054-3
Shattuck DW, Sandor-Leahy SR, Schaper KA, Rottenberg DA, Leahy RM (2001) Magnetic resonance image tissue classification using a partial volume model. Neuroimage 13(5):856–876. https://doi.org/10.1006/nimg.2000.0730
Joshi, A. A., Bhushan, C., Salloum, R., Wisnowski, J. L., Shattuck, D. W., & Leahy, R. M. (2018). Using the anisotropic laplace equation to compute cortical thickness. Medical Image Computing and Computer Assisted Intervention – MICCAI 2018, Cham.
Joshi, A. A., Shattuck, D. W., & Leahy, R. M. (2012). A method for automated cortical surface registration and labeling. Biomedical Image Registration, Berlin, Heidelberg.
Joshi AA, Shattuck DW, Thompson PM, Leahy RM (2007) Surface-constrained volumetric brain registration using harmonic mappings. IEEE Trans Med Imaging 26(12):1657–1669. https://doi.org/10.1109/TMI.2007.901432
Joshi AA, Choi S, Liu Y, Chong M, Sonkar G, Gonzalez-Martinez J, Nair D, Wisnowski JL, Haldar JP, Shattuck DW, Damasio H, Leahy RM (2022) A hybrid high-resolution anatomical MRI atlas with sub-parcellation of cortical gyri using resting fMRI. J Neurosci Methods 374:109566. https://doi.org/10.1016/j.jneumeth.2022.109566
Tierney TM, Alexander NA, Ashburner J, Avila NL, Balbastre Y, Barnes G, Bezsudnova Y, Brudfors M, Eckstein K, Flandin G, Friston K, Jafarian A, Kowalczyk OS, Litvak V, Medrano J, Mellor S, O’Neill G, Parr T, Razi A, Zeidman P (2025) SPM 25: open source neuroimaging analysis software. J Open Source Softw 10(110):8103. https://doi.org/10.21105/joss.08103
Manjón JV, Coupé P, Martí-Bonmatí L, Collins DL, Robles M (2010) Adaptive non-local means denoising of MR images with spatially varying noise levels. J Magn Reson Imaging 31(1):192–203. https://doi.org/10.1002/jmri.22003
Ashburner J, Friston KJ (2005) Unified segmentation. Neuroimage 26(3):839–851. https://doi.org/10.1016/j.neuroimage.2005.02.018
Rajapakse JC, Giedd JN, Rapoport JL (1997) Statistical approach to segmentation of single-channel cerebral MR images. IEEE Trans Med Imaging 16(2):176–186. https://doi.org/10.1109/42.563663
Ashburner J, Friston KJ (2011) Diffeomorphic registration using geodesic shooting and Gauss–Newton optimisation. Neuroimage 55(3):954–967. https://doi.org/10.1016/j.neuroimage.2010.12.049
Keshavan A, Yeatman JD, Rokem A (2019) Combining citizen science and deep learning to amplify expertise in neuroimaging. Front Neuroinform. https://doi.org/10.3389/fninf.2019.00029
JASP Team. (2025). JASP (Version 0.95.1). In https://jasp-stats.org/
Cosgrove KP, Mazure CM, Staley JK (2007) Evolving knowledge of sex differences in brain structure, function, and chemistry. Biol Psychiatry 62(8):847–855. https://doi.org/10.1016/j.biopsych.2007.03.001
Ruigrok ANV, Salimi-Khorshidi G, Lai M-C, Baron-Cohen S, Lombardo MV, Tait RJ, Suckling J (2014) A meta-analysis of sex differences in human brain structure. Neurosci Biobehav Rev 39:34–50. https://doi.org/10.1016/j.neubiorev.2013.12.004
Backhausen LL, Herting MM, Buse J, Roessner V, Smolka MN, Vetter NC (2016) Quality control of structural MRI images applied using FreeSurfer—a hands-on workflow to rate motion artifacts. Front Neurosci 10:558. https://doi.org/10.3389/fnins.2016.00558
Cohen J (1988) Statistical power analysis for the behavioral sciences. Routledge Academic
Esteban O, Blair RW, Nielson DM, Varada JC, Marrett S, Thomas AG, Poldrack RA, Gorgolewski KJ (2019) Crowdsourced MRI quality metrics and expert quality annotations for training of humans and machines. Sci Data 6:30. https://doi.org/10.1038/s41597-019-0035-4
Schirmer MD, Dalca AV, Sridharan R, Giese A-K, Donahue KL, Nardin MJ, Mocking SJT, McIntosh EC, Frid P, Wasselius J, Cole JW, Holmegaard L, Jern C, Jimenez-Conde J, Lemmens R, Lindgren AG, Meschia JF, Roquer J, Rundek T, Rost NS (2019) White matter hyperintensity quantification in large-scale clinical acute ischemic stroke cohorts – the MRI-GENIE study. Neuroimage Clin 23:101884. https://doi.org/10.1016/j.nicl.2019.101884
White T, Jansen PR, Muetzel RL, Sudre G, El Marroun H, Tiemeier H, Qiu A, Shaw P, Michael AM, Verhulst FC (2018) Automated quality assessment of structural magnetic resonance images in children: comparison with visual inspection and surface-based reconstruction. Hum Brain Mapp 39(3):1218–1231. https://doi.org/10.1002/hbm.23911
Seidler RD, Bernard JA, Burutolu TB, Fling BW, Gordon MT, Gwin JT, Kwak Y, Lipps DB (2010) Motor control and aging: links to age-related brain structural, functional, and biochemical effects. Neurosci Biobehav Rev 34(5):721–733. https://doi.org/10.1016/j.neubiorev.2009.10.005
Botvinik-Nezer R, Wager TD (2023) Reproducibility in neuroimaging analysis: challenges and solutions. Biol Psychiatry Cogn Neurosci Neuroimaging 8(8):780–788. https://doi.org/10.1016/j.bpsc.2022.12.006
Appelbaum M, Cooper H, Kline RB, Mayo-Wilson E, Nezu AM, Rao SM (2018) Journal article reporting standards for quantitative research in psychology: the APA publications and communications board task force report. Am Psychol 73(1):3–25. https://doi.org/10.1037/amp0000191
Acknowledgements
The author would like to thank Huang Xindi for offering advice, encouragement, and assistance in the completion of this work.
Funding
The Healthy Brain Network (http://www.healthybrainnetwork.org) and its collaborative initiatives are supported by philanthropic contributions from the following individuals, foundations and organizations: Margaret Bilotti; Brooklyn Nets; Agapi and Bruce Burkard; James Chang; Phyllis Green and Randolph Cowen; Grieve Family Fund; Susan Miller and Byron Grote; Sarah and Geoff Gund; George Hall; Jonathan M. Harris Family Foundation; Joseph P. Healey; The Hearst Foundations; Eve and Ross Jaffe; Howard & Irene Levine Family Foundation; Rachael and Marshall Levine; George and Nitzia Logothetis; Christine and Richard Mack; Julie Minskoff; Valerie Mnuchin; Morgan Stanley Foundation; Amy and John Phelan; Roberts Family Foundation; Jim and Linda Robinson Foundation, Inc.; Linda and Richard Schaps; Zibby Schwarzman; Abigail Pogrebin and David Shapiro; Stavros Niarchos Foundation; Preethi Krishna and Ram Sundaram; Amy and John Weinberg; Donors to the 2013 Child Advocacy Award Dinner Auction; Donors to the 2012 Brant Art Auction.
Author information
Authors and Affiliations
Contributions
YSW was responsible for conceptualization, methodology, formal analysis, investigation, data curation, validation, visualization, and writing (original draft, review, and editing).
Corresponding author
Ethics declarations
Ethics approval
All procedures performed were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Chesapeake Institutional Review Board (now Advarra, Inc.; https://www.advarra.com/; protocol number Pro00012309).
Consent to participate
All participants provided written assent and/or written informed consent, including consent to share de-identified data.
Consent for publication
The author affirms that human research participants provided informed consent for publication.
Conflict of interest
The author declares no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wong, YS. Validation of five automated structural MRI quality assessment tools against expert ratings. Neuroradiology (2026). https://doi.org/10.1007/s00234-026-03942-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1007/s00234-026-03942-9